Digimouse atlas FEM mesh
Version 1
- Version 1 was created by Qianqian Fang <fangq at nmr.mgh.harvard.edu>, released on 03/31/2011
- License: This mesh is released in the public domain, feel free to use for any purposes
- 1. Download
- 2. Introduction
- 3. Format
- 4. How is this mesh different from Digimouse tessellated atlas
- 5. What's in the package
- 6. Tissue optical properties
- 7. How to recreate this mesh
- 8. Footnote
- 9. Reference
1. Download
Please download the latest release (Version 1) at our registration/download page.
2. Introduction
Digimouse is a popular mouse atlas (Dogdas2007). FEM mesh Version 1 was created by Qianqian Fang using iso2mesh (Fang2009) version 1.0 and CGAL (CGAL2009).
URL: http://mcx.sourceforge.net/cgi-bin/index.cgi?MMC/DigimouseMesh
3. Format
The mesh data were saved in matlab/octave .mat format. Please load the .mat file in matlab/octave and export to your FEM simulator.
node: node coordinates (in mm) face: surface triangles; the last column is the surface ID, elem: tetrahedral elements; the last column is the region ID
The low-resolution mesh, (1L), has 42301 nodes, 210161 tetrahedral elements, 149158 triangular surface elements and 21 tissue segmentations.
4. How is this mesh different from Digimouse tessellated atlas
This mesh has several merits compared to the tessellated atlas provided at Digimouse's website:
- it contains 30% less nodes, but better spatial sampling and surface smoothness
- it contains all 21 tissue segmentations, while the tessellated atlas has 17 of them
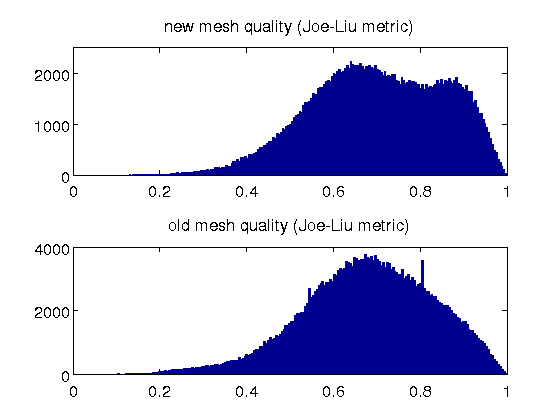
- the mesh quality is slightly improved, with an average Joe-Liu metric of 0.7; that for the original mesh is 0.68
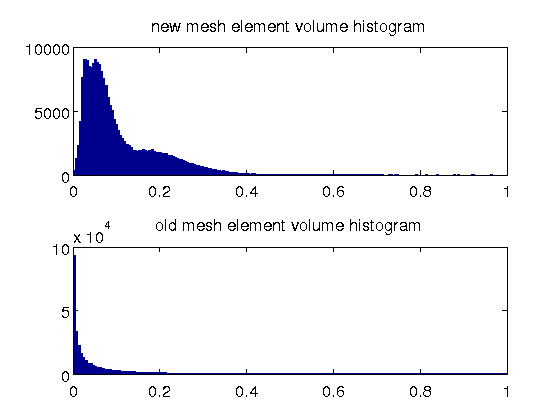
- the element size in the new mesh has a much better spectrum; there are a large number of small elements in the old mesh
5. What's in the package
- digimouse_tissue_index.txt
- tissue index for the original atlas segmentation this should match the last column of elem
- digimouse_optical_properties.dat
- tissue optical properties, in the order of [mua (1/mm), mus (1/mm), anisotropy, refractive index]
- digimouse_mesh.m
- mesh generation script to create this mesh using iso2mesh 1.0
6. Tissue optical properties
The optical properties of different tissue types are listed below. The values are largely based on two papers: (Cheong1990) and (Strangman2003). Please see the notes in the table below.
| Tissue id | Tissue type | μa (mm−1) | μs (mm−1) | Anisotropy (g) | Refract. Index (n) | Notes |
|---|---|---|---|---|---|---|
| 1 | skin | 0.0191 | 6.6 | 0.9 | 1.37 | skin here is actually soft tissue, use scalp data @830nm from Strangman2003 |
| 2 | skeleton | 0.0136 | 8.6 | 0.9 | 1.37 | use skull from Strangman2003 |
| 3 | eye | 0.0026 | 0.01 | 0.9 | 1.37 | use CSF from Strangman2003 |
| 4 | medulla | 0.0186 | 11.1 | 0.9 | 1.37 | use brain from Strangman2003 |
| 5 | cerebellum | 0.0186 | 11.1 | 0.9 | 1.37 | brain |
| 6 | olfactory bulbs | 0.0186 | 11.1 | 0.9 | 1.37 | brain |
| 7 | external cerebrum | 0.0186 | 11.1 | 0.9 | 1.37 | brain |
| 8 | striatum | 0.0186 | 11.1 | 0.9 | 1.37 | brain |
| 9 | heart | 0.0240 | 8.9 | 0.9 | 1.37 | muscle |
| 10 | rest of the brain | 0.0026 | 0.01 | 0.9 | 1.37 | use CSF from Strangman2003 |
| 11 | masseter muscles | 0.0240 | 8.9 | 0.9 | 1.37 | muscle |
| 12 | lachrymal glands | 0.0240 | 8.9 | 0.9 | 1.37 | muscle |
| 13 | bladder | 0.0240 | 8.9 | 0.9 | 1.37 | muscle |
| 14 | testis | 0.0240 | 8.9 | 0.9 | 1.37 | muscle |
| 15 | stomach | 0.0240 | 8.9 | 0.9 | 1.37 | muscle |
| 16 | spleen | 0.072 | 5.6 | 0.9 | 1.37 | use rat liver data from Cheong1990 |
| 17 | pancreas | 0.072 | 5.6 | 0.9 | 1.37 | use rat liver data from Cheong1990 |
| 18 | liver | 0.072 | 5.6 | 0.9 | 1.37 | from Cheong1990 |
| 19 | kidneys | 0.050 | 5.4 | 0.9 | 1.37 | use cow kidney @789nm from Cheong1990 |
| 20 | adrenal glands | 0.024 | 8.9 | 0.9 | 1.37 | muscle |
| 21 | lungs | 0.076 | 10.9 | 0.9 | 1.37 | use pig lung @850nm from Cheong1990, change g to 0.9 |
7. How to recreate this mesh
A matlab/octave script, digimouse_mesh.m, is provided in the downloaded mesh package to allow you to recreate this mesh. In addition, you also need to download the segmented Digimouse atlas Analyze data from it's website and saved in the same folder as the script. You also to download and configure iso2mesh 1.0 or above to run this script.
The mesh generation script is listed below:
%% load the segmented Digimouse mouse atlas
fid=fopen('atlas_380x992x208.img','rb');
digimouse=fread(fid,inf,'uchar');
fclose(fid);
dim=[380 992 208];
digimouse=uint8(reshape(digimouse,dim));
%% perform mesh generation using iso2mesh and CGAL 3D mesher
opt.radbound=5; % max surface triangle diameter
opt.distbound=3; % related to the distance to the tissue boundary
[node,elem,face]=v2m(digimouse,[],opt,10,'cgalmesh');
%% post processing, scale mesh with the voxel size 0.1 mm
node(:,4)=[];
node=node*0.1;
elem(:,1:4)=meshreorient(node(:,1:3),elem(:,1:4));
%% packaging the data
README_digimouse_mesh= ...
sprintf(['Digimouse atlas FEM mesh - Version 1L (low-resolution).\n\n' ...
'Created on 02/11/2011 by Qianqian Fang [1] with iso2mesh version 1.0.0 [2]\n' ...
'and CGAL [3]. The image segmentation was downloaded from [4].\n\n' ...
'License: \n\tpublic domain\n' ...
'Format: \n' ...
' node: node coordinates (in mm)\n' ...
' face: surface triangles, the last column is the tissue surface ID \n' ...
' elem: tetrahedral elements, the last column is the region ID \n\n' ...
'Other files:\n' ...
' digimouse_tissue_index.txt: tissue index for the original atlas segmentation\n' ...
' this should match the last column of elem\n' ...
' digimouse_optical_properties.dat: tissue optical properties, in the order of\n' ...
' [mua (1/mm), mus (1/mm), anisotropy, refractive index] \n' ...
' digimouse_mesh.m: mesh generation script to create this mesh using iso2mesh 1.0\n' ...
'URL: http://mcx.sourceforge.net/cgi-bin/index.cgi?MMC/DigimouseMesh\n\n' ...
'Links:\n [1] http://nmr.mgh.harvard.edu/~fangq/\n [2] http://iso2mesh.sf.net\n' ...
' [3] http://www.cgal.org/\n [4] http://neuroimage.usc.edu/Digimouse_download.html\n\n' ...
'Related Citations\n' ...
' [iso2mesh]: Q. Fang and D. Boas, "Tetrahedral mesh generation from volumetric binary and gray-scale images," \n' ...
' Proceedings of IEEE International Symposium on Biomedical Imaging 2009, pp. 1142-1145, 2009\n' ...
' [CGAL]: L. Rineau, S. Tayeb, and M. Yvinec, "3D Mesh Generation," In CGAL Editorial Board, \n' ...
' Ed, CGAL User and Reference Manual. 3.5 edition, 2009\n' ...
' [Digimouse]:B. Dogdas, D. Stout, A. Chatziioannou, and R. M. Leahy, "Digimouse: A 3D Whole Body Mouse \n' ...
' Atlas from CT and Cryosection Data." Phys Med Biol. 52 (3): 577-87, 2007\n']);
save Digimouse_Mesh_1L.mat node elem face README_digimouse_mesh
8. Footnote
The mesh data is in the public domain.
If you use this mesh in your publication, please cite the mesh version number to avoid conflict to any further updates of this mesh. We are also appreciated if you can cite the papers for iso2mesh or CGAL in your publication.
9. Reference
[Fang2009] Q. Fang and D. Boas, “Tetrahedral mesh generation from volumetric binary and gray-scale images,” Proceedings of IEEE International Symposium on Biomedical Imaging 2009, 1142–1145 (2009).
[CGAL2009] L. Rineau, S. Tayeb, and M. Yvinec. 3D Mesh Generation. In CGAL Editorial Board, Ed, CGAL User and Reference Manual. 3.5 edition, 2009
[Dogdas2007] B. Dogdas, D. Stout, A. Chatziioannou, and R. M. Leahy, "Digimouse: A 3D Whole Body Mouse Atlas from CT and Cryosection Data." Phys Med Biol. 52 (3): 577-87, 2007
[Cheong1990] W. F. Cheong, S. A. Prahl, and A. J. Welch, "A Review of the Optical Properties of Biological Tissues," IEEE J. Quantum Electronics, 26, 2166-2185, 1990.
[Strangman2003] G. Strangman, M. A. Franceschini and D. A. Boas. Factors affecting the accuracy of near-infrared spectroscopy concentration calculations for focal changes in oxygenation parameters. Neuroimage, 18(4), 865-879, 2003
![[Home]](/img/mcx_wiki_banner.png)